Data + Coordinate Injection¶
This page explains how location and date features are fused and injected into the denoiser using one FiLM-style conditioning path.
Where This Happens in Code¶
- coordinate/date encoding logic:
src/depth_recon/models/diffusion/DenoisingDiffusionProcess/DenoisingDiffusionProcess.py - FiLM application in ConvNeXt block:
src/depth_recon/models/diffusion/DenoisingDiffusionProcess/backbones/unet_convnext.py
Enabling It¶
Main config flags (model section):
- coord_conditioning.enabled
- coord_conditioning.encoding: unit_sphere, sincos, raw
- coord_conditioning.include_date
- coord_conditioning.date_encoding: currently day_of_year_sincos
- coord_conditioning.embed_dim
Data-side requirement:
- dataset.output.return_coords: true so batches include coords
Runtime requirements enforced by code:
- if coord conditioning is enabled and coords is missing, inference/training raises an error
- if date conditioning is enabled and date is missing, inference/training raises an error
Coordinate Encoding Options¶
unit_sphere (default)¶
Input: latitude/longitude in degrees.
Transform:
- convert to radians
- map to 3D unit sphere
- output features: (x, y, z)
Why it is useful:
- avoids longitude discontinuity at ±180°
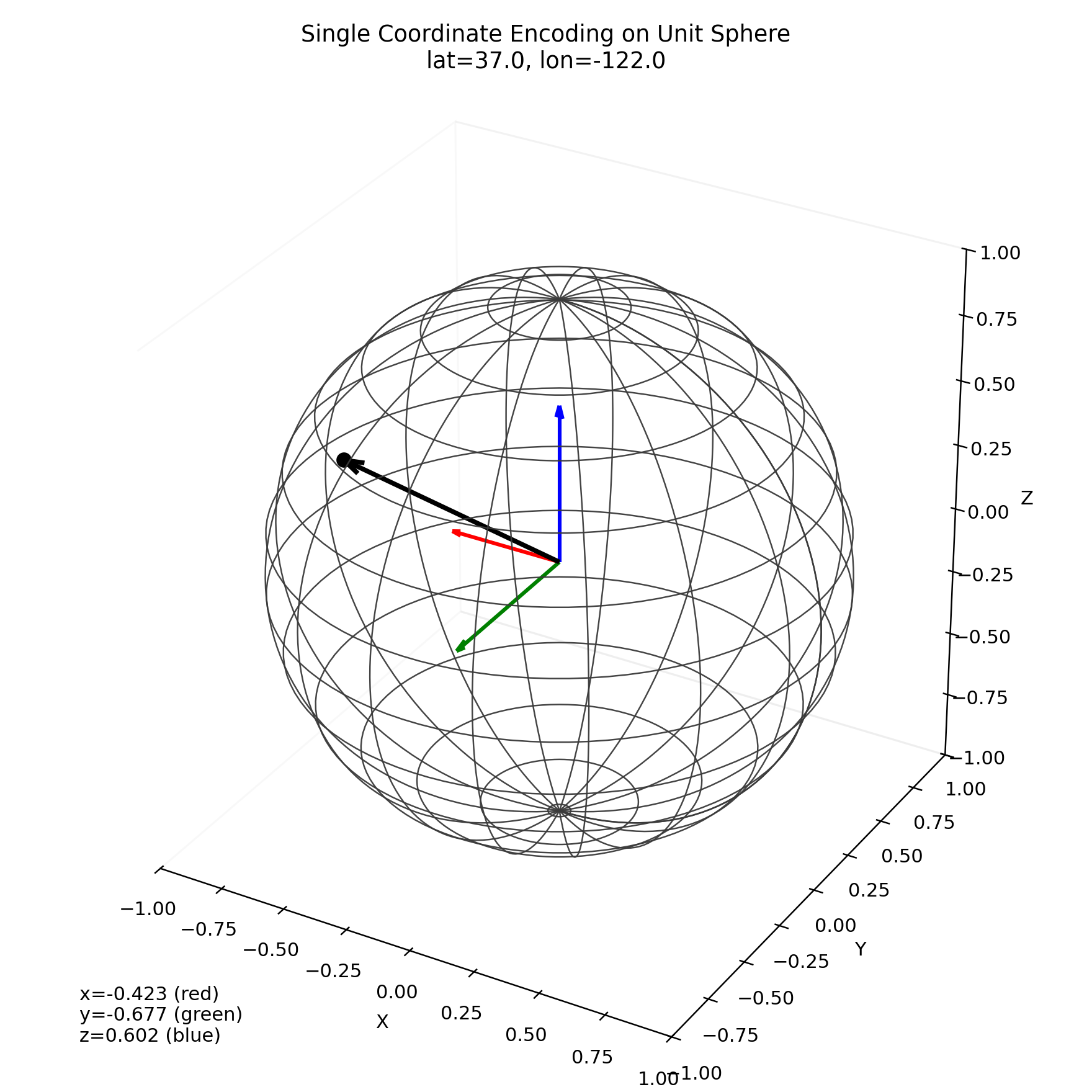
This 3D view shows one example coordinate encoded as (x, y, z) on the sphere. Red/green/blue arrows are the x/y/z components, and the black arrow is the resulting encoded vector.
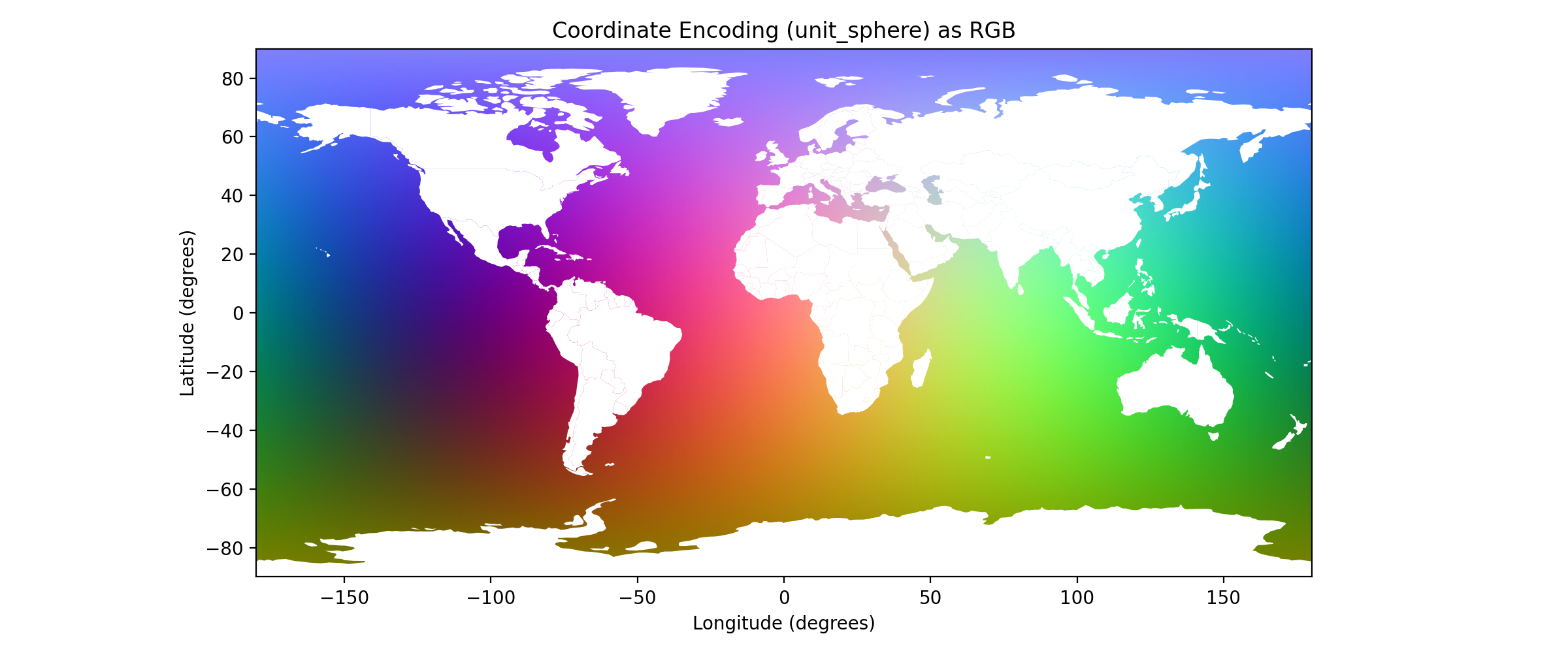
This image evaluates all integer (lat, lon) combinations and maps the 3D unit-sphere encoding (x, y, z) to RGB. It visualizes how each coordinate is transformed into a 3-value vector before FiLM embedding, showing smooth transitions and proper wrap-around from -90 to +90 and -180 to +180.
sincos¶
Features:
- sin(lat), cos(lat), sin(lon), cos(lon)
Why it is useful:
- periodic and wrap-safe representation
raw¶
Features:
- lat / 90
- lon / 180
Why it is simple:
- minimal feature transform, but not wrap-safe at dateline transitions
Date Encoding¶
Current implementation supports one option:
- day_of_year_sincos
Pipeline:
1. parse date as integer YYYYMMDD
2. validate month/day values
3. compute non-leap day-of-year using fixed month offsets
4. encode as:
- sin(2*pi*doy/365)
- cos(2*pi*doy/365)
Dataset date parsing convention:
- monthly source names (YYYYMM) are converted to mid-month (YYYYMM15)
- in the OSTIA overlap setup, depth files still provide source_file while OSTIA timestamps are stored separately (ostia_timestamp_utc), so model date conditioning remains mid-month aligned
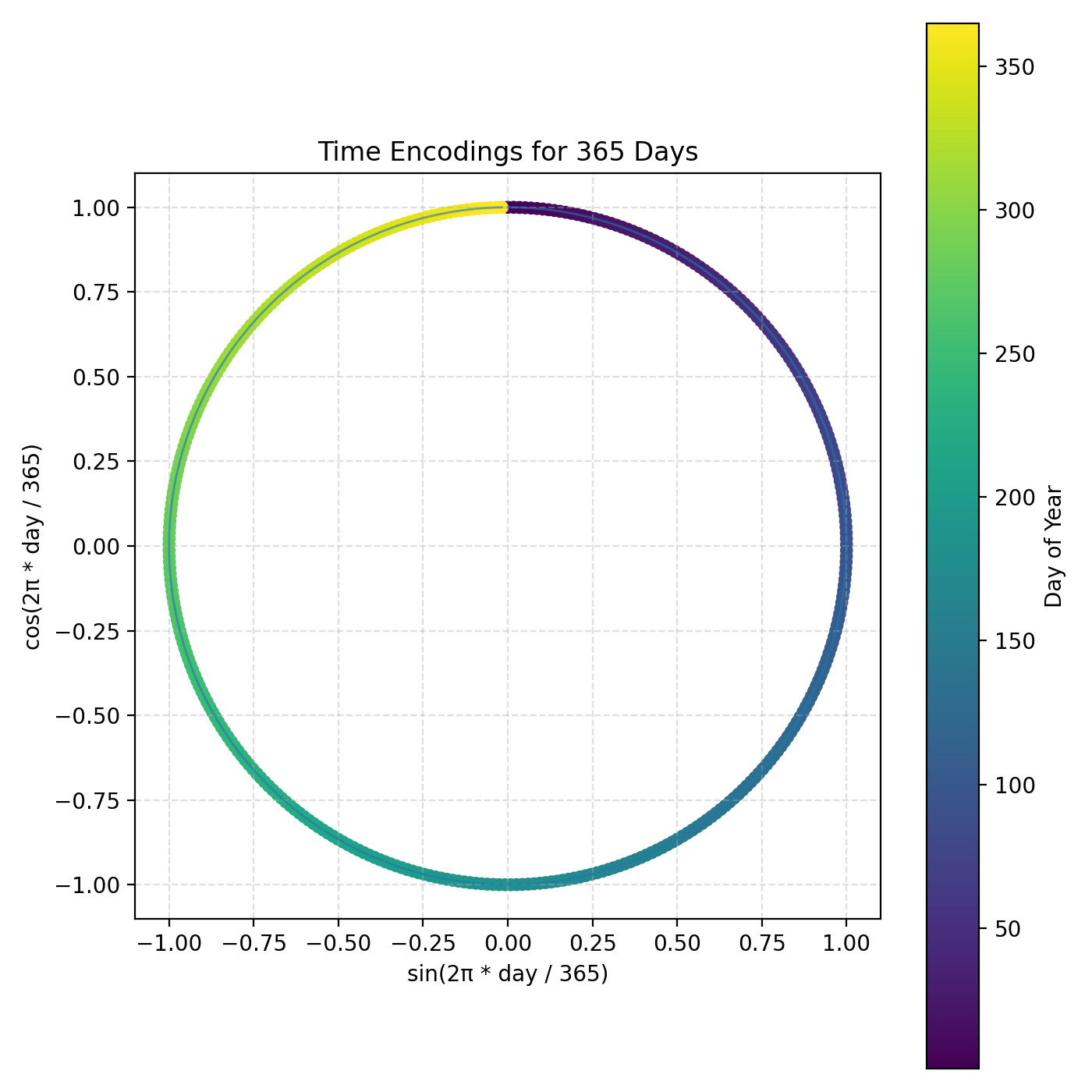
This plot shows all 365 day-of-year embeddings as points in the sin/cos plane. Each day maps to one point on the unit circle, which makes the representation periodic and year-wrap safe.
Feature Fusion¶
When include_date=true:
- encoded date features are concatenated with encoded coordinate features
- the concatenated vector is passed through a small MLP (coord_mlp) to produce one joint conditioning embedding
FiLM Injection Mechanism¶
Exact Injection Path (Code)¶
- embedding creation:
DenoisingDiffusionConditionalProcess._maybe_embed_coords(...)insrc/depth_recon/models/diffusion/DenoisingDiffusionProcess/DenoisingDiffusionProcess.py - U-Net call site:
DenoisingDiffusionConditionalProcess.forward(...)andp_loss(...)passcoord_embtoself.model(..., coord_emb=coord_emb) - injection site:
ConvNextBlock.forward(...)insrc/depth_recon/models/diffusion/DenoisingDiffusionProcess/backbones/unet_convnext.py
End-to-End Pseudocode (What Happens, In Order)¶
# in DenoisingDiffusionConditionalProcess.forward(...) and p_loss(...)
coord_emb = None
if coord_conditioning_enabled:
# 1) Encode lat/lon -> encoded_coord (unit_sphere | sincos | raw)
encoded_coord = _encode_coords(coord) # shape: (B, coord_feat_dim)
# 2) Optionally encode date and concatenate
if date_conditioning_enabled:
encoded_date = _encode_date(date) # shape: (B, 2) for day_of_year_sincos
encoded_coord = concat(encoded_coord, encoded_date, dim=1)
# 3) Project to one shared conditioning vector used by all blocks
# coord_mlp: Linear(enc_dim -> 4*E) -> GELU -> Linear(4*E -> E)
coord_emb = coord_mlp(encoded_coord) # shape: (B, E)
# reverse/training pass
prediction = unet(model_input, t, coord_emb=coord_emb)
# in UnetConvNextBlock.forward(...)
t_emb = time_mlp(t) # diffusion-step embedding (if enabled)
# pass the same coord_emb to every ConvNext block in down, mid, up, and final block
for each ConvNextBlock in model:
x = block(x, time_emb=t_emb, coord_emb=coord_emb)
# in ConvNextBlock.forward(...)
h = depthwise_conv7x7(x)
if time_emb is available:
# additive timestep conditioning
h = h + time_mlp_block(time_emb)[:, :, None, None]
if coord_emb is available:
# FiLM parameters from shared coord/date embedding
# block-specific coord_mlp: GELU -> Linear(E -> 2*C)
scale_shift = coord_mlp_block(coord_emb) # shape: (B, 2*C)
scale, shift = split(scale_shift, 2, dim=1) # each: (B, C)
h = h * (1 + scale[:, :, None, None]) + shift[:, :, None, None]
# then regular ConvNeXt transform + residual
h = convnext_subnet(h)
out = h + residual_projection(x)
What This Means Practically¶
- coord/date are fused once into a single
coord_emb, then reused everywhere in the U-Net - FiLM is applied per sample and per channel, with spatial broadcasting over
H x W - injection is multiplicative + additive (
1 + scale,shift), so near-zero FiLM output behaves close to identity - timestep context and geo/date context are both active in each conditioned block, but with different operations (additive time, FiLM coord/date)